Microbiological characterization of raw goat milk from a small municipality in Brazil: identification of Gram-positive cocci by MALDI-TOF MS
DOI:
https://doi.org/10.70099/BJ/2026.03.02.1Keywords:
Staphylococcus aureus, dairy, mass spectrometryAbstract
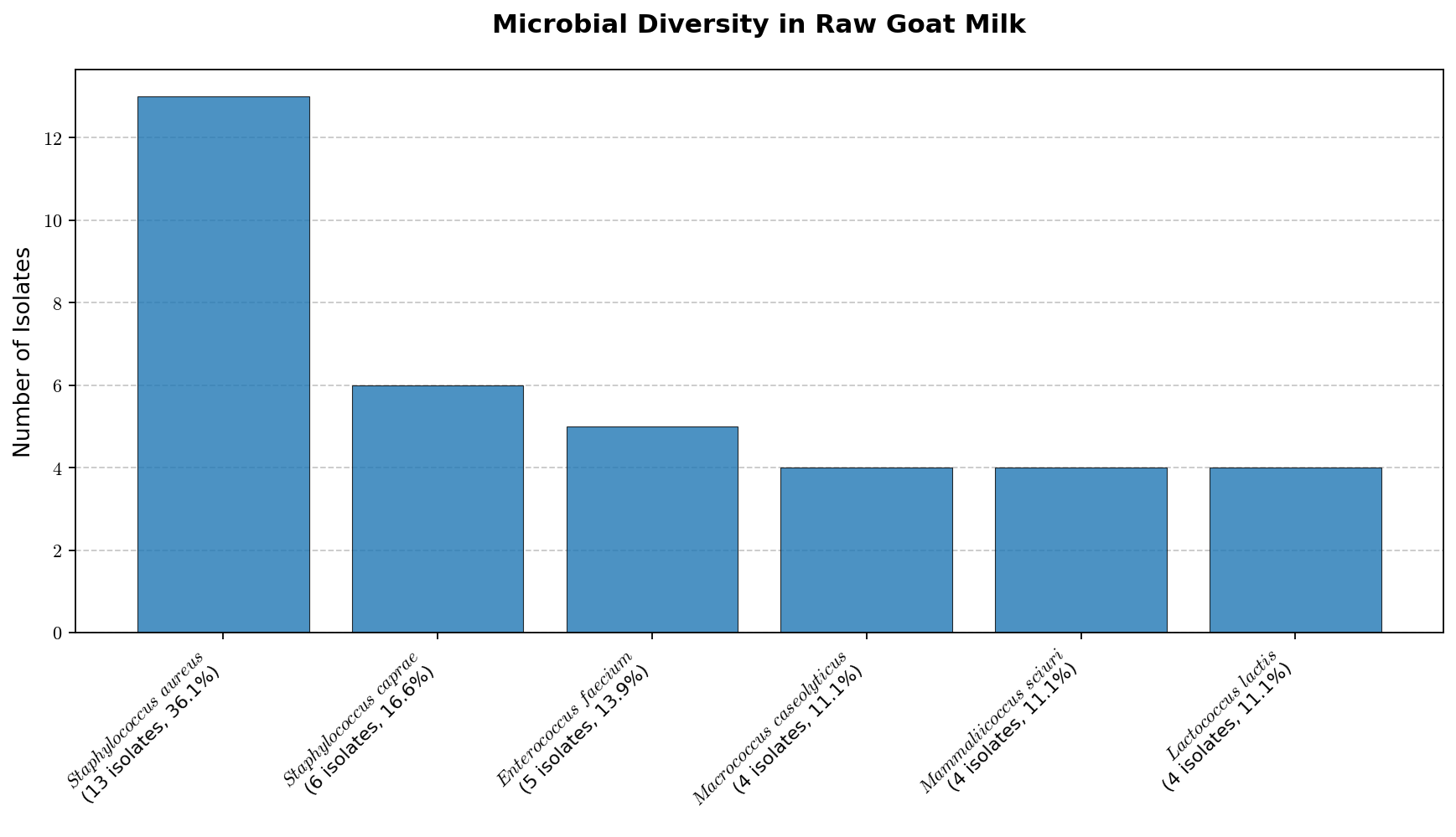
Goat milk is a high-nutritional-value food, but it can also harbor several pathogenic microorganisms. It is not uncommon for this food to be consumed raw, which can pose health risks to consumers. In this study, we sought to verify the diversity of Gram-positive cocci (GPC), especially Staphylococcus sp., in raw milk sam-ples obtained in the region of Itaocara, RJ. Microbial identification of the nine samples was performed by mass spectrometry (MALDI-TOF) after isolation in selective culture media. MALDI-TOF analysis of the isolates from raw goat milk samples revealed the predominance of Staphylococcus spp., with emphasis on S. aureus and S. caprae. GPCs of other genera were also detected. Surprisingly, some of these microorganisms were isolated on selective media for Gram-negative bacteria, highlighting the importance of accurate confirm-atory identification methods such as MALDI-TOF. The detection of S. aureus, a relevant foodborne pathogen, and other species with spoilage or pathogenic potential, such as S. caprae and Mammaliicoccus sciuri, in six of the nine samples highlights potential microbiological risks associated with locally consumed raw goat milk. On the other hand, the presence of lactic acid bacteria (Lactococcus lactis, L. garvieae, and Enterococcus sp.), with technological potential for producing dairy products, was also observed. The recovery of Gram-positive cocci from media selective for Gram-negative bacteria reinforces the limitations of culture-based selectivity and highlights the importance of confirmatory identification methods such as MALDI-TOF MS. Keywords: Staphylococcus aureus, dairy product, mass spectrometry
References
1. Moraes MS, Ramos GLP, Nascimento JS (2025) Benefits of goat, sheep and buffalo milk: A concise review. Int J Vet Sci Anim Husb 10:150–155.
2. Roy D, Ye A, Moughan PJ, Singh H (2020) Composition, structure, and digestive dynamics of milk from different species—A review. Front Nutr 7:577759. https://doi.org/10.3389/fnut.2020.577759
3. Fusco V, Chieffi D, Fanelli F, Logrieco AF, Cho GS, Kabisch J, Arioli F, Papademas P, De Angelis M, Franz CM (2020) Microbial quality and safety of milk and milk products in the 21st century. Compr Rev Food Sci Food Saf 19:2013–2049. https://doi.org/10.1111/1541-4337.12568
4. Ruiz P, Barragán I, Seseña S, Palop ML (2016) Is staphylococci population from milk of healthy goats safe? Int J Food Microbiol 238:146–152. https://doi.org/10.1016/j.ijfoodmicro.2016.09.006
5. Wiśniewski P, Gajewska J, Zadernowska A, Chajęcka-Wierzchowska W (2024) Identification of the enterotoxigenic potential of Staphylococcus spp. from raw milk and raw milk cheeses. Toxins 16:17. https://doi.org/10.3390/toxins16010017
6. Machado MAA, Ribeiro WA, Toledo VS, Ramos GLPA, Vigoder HC, Nascimento JS (2020) Anti-biotic resistance and biofilm production in Gram-positive catalase-positive cocci isolated from Brazilian pasteurized milk. J Food Qual Hazards Control 7:67–74.
7. Ramos GLPA, Vigoder HC, Nascimento JS (2021) Technological applications of Macrococcus case-olyticus and its impact on food safety. Curr Microbiol 78:11–16. https://doi.org/10.1007/s00284-020-02264-5
8. Ruiz-Romero RA, Vargas-Bello-Pérez E (2023) Non-aureus staphylococci and mammaliicocci as a cause of mastitis in domestic ruminants: A systematic review. Vet Res Commun 47:1067–1084. https://doi.org/10.1007/s11259-023-10135-0
9. Terzić-Vidojević A, Veljović K, Popović N, Tolinački M, Golić N (2021) Enterococci from raw-milk cheeses: Current knowledge on safety, technological, and probiotic concerns. Foods 10:2753. https://doi.org/10.3390/foods10112753
10. International Organization for Standardization (1999) ISO 6888-1:1999. Microbiology of food and animal feeding stuffs—Horizontal method for the enumeration of coagulase-positive staphylococci (Staphylococcus aureus and other species)—Part 1: Technique using Baird-Parker agar medium.
11. Bonnet M, Lagier JC, Raoult D, Khelaifia S (2019) Bacterial culture through selective and non-selective conditions: the evolution of culture media in clinical microbiology. New Microbes New Infect 34:100622. https://doi.org/10.1016/j.nmni.2019.100622
12. Calderaro A, Chezzi C (2024) MALDI-TOF MS: A reliable tool in the real life of the clinical micro-biology laboratory. Microorganisms 12:322. https://doi.org/10.3390/microorganisms12020322
13. Rychert J (2019) Benefits and limitations of MALDI-TOF mass spectrometry for the identification of microorganisms. J Infectiol Epidemiol 2:1510109. https://doi.org/10.23937/2474-3658/1510109
14. Grispoldi L, Karama M, Armani A, Hadjicharalambous C, Cenci-Goga BT (2021) Staphylococcus aureus enterotoxin in food of animal origin and staphylococcal food poisoning risk assessment from farm to table. Ital J Anim Sci 20:677–690. https://doi.org/10.1080/1828051X.2021.1901263
15. Zhang J, Wang J, Jin J, Li X, Zhang H, Shi X, Zhao C (2022) Prevalence, antibiotic resistance, and enterotoxin genes of Staphylococcus aureus isolated from milk and dairy products worldwide: A systematic review and meta-analysis. Food Res Int 162:111969. https://doi.org/10.1016/j.foodres.2022.111969
16. Nelli A, Voidarou C, Venardou B, Fotou K, Tsinas A, Bonos E, Fthenakis GC, Skoufos I, Tzora A (2022) Antimicrobial and methicillin resistance pattern of potential mastitis-inducing Staphylococcus aureus and coagulase-negative staphylococci isolates from dairy goats. Biology 11:1591. https://doi.org/10.3390/biology11111591
17. Mastroianni A, Greco S, Vangeli V, Mauro MV, Manfredi R (2023) Invasive diseases due to fatal Staphylococcus caprae infection: A case series and literature review. J Infect Dis Epidemiol 9:298. https://doi.org/10.23937/2474-3658/1510298
18. Mazhar S, Kilcawley KN, Hill C, McAuliffe O (2020) A systems-wide analysis of proteolytic and lipolytic pathways uncovers the flavor-forming potential of the Gram-positive bacterium Macrococcus caseolyticus subsp. caseolyticus. Front Microbiol 11:1533. https://doi.org/10.3389/fmicb.2020.01533
19. Madhaiyan M, Wirth JS, Saravanan VS (2020) Phylogenomic analyses of the Staphylococcaceae family suggest taxonomic reclassification within the genus Staphylococcus. Int J Syst Evol Microbiol 70:5926–5936. https://doi.org/10.1099/ijsem.0.004498
20. Crippa BL, Matos LG, Souza FN, Silva NCC (2024) Non-aureus staphylococci and mammaliicocci (NASM): their role in bovine mastitis and One health. J Dairy Res 91:1–13. https://doi.org/10.1017/S0022029924000165
21. Fatahi-Bafghi M (2021) Characterization of the Rothia spp. and their role in human clinical infections. Infect Genet Evol 88:104686. https://doi.org/10.1016/j.meegid.2020.104686
22. Ruaro A, Andrighetto C, Torriani S, Lombardi A (2013) Biodiversity and characterization of indig-enous coagulase-negative staphylococci isolated from raw milk and cheese of North Italy. Food Mi-crobiol 34:106–111. https://doi.org/10.1016/j.fm.2012.11.017
23. Saiki R, Hagi T, Narita T, Kobayashi M, Sasaki K, Asahina Y, Nomura M (2018) Effects of the ad-dition of non-starter lactic acid bacteria on free amino acid production during cheese ripening. Food Sci Technol Res 24:299–309.
24. Kondrotiene K, Zavistanaviciute P, Aksomaitiene J, Novoslavskij A, Malakauskas M (2024) Lacto-coccus lactis in dairy fermentation—Health-promoting and probiotic properties. Fermentation 10:16. https://doi.org/10.3390/fermentation10010016
25. Perin LM, Belviso S, Dal Bello B, Nero LA, Cocolin L (2017) Technological properties and biogenic amines production by bacteriocinogenic lactococci and enterococci strains isolated from raw goat's milk. J Food Prot 80:151–157. https://doi.org/10.4315/0362-028X.JFP-16-255
Published
How to Cite
License
Copyright (c) 2026 Gustavo Luis de Paiva Anciens Ramos, Emily Cristine Candido da Silva, Lavínia Falcão de Araújo, Marcelo Soares de Moraes, Janaína dos Santos Nascimento (Author)

This work is licensed under a Creative Commons Attribution 4.0 International License.
Authors who publish with BioNatura Journal agree to the following terms: Authors retain copyright and grant the BioNatura Institutional Publishing Consortium (BIPC) right of first publication with the work simultaneously licensed under a Creative Commons Attribution 4.0 International License (CC BY 4.0). This allows others to share the work with an acknowledgment of the work's authorship and initial publication in this journal.This is an open access article distributed under the terms of the Creative Commons Attribution 4.0 International License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.




