MINI-REVIEW
Role of Probiotics in Gut Health: Mechanisms, Clinical Relevance, And Translational Challenges – A Critical Comparative Review
ABSTRACT
Probiotics have been extensively investigated as modulators of the gut microbiota and host physiology; however, clinical efficacy remains inconsistent despite substantial mechanistic evidence. This review critically evaluates the gap between mechanistic findings and clinical outcomes using a comparative analytical framework. Probiotic efficacy is highly dependent on strain specificity, host microbiome composition, dosage, and study design. Mechanisms—including competitive exclusion, antimicrobial metabolite production (short-chain fatty acids, indoles, secondary bile acids), immunomodulation (NF-κB inhibition, IL-10 induction), and gut barrier enhancement (tight junction regulation via AMPK and occludin/claudin expression)—are strongly supported by experimental evidence. However, clinical translation is limited by host-specific colonization resistance, baseline microbiota composition, dietary substrate availability, and methodological heterogeneity across trials. A multilayer interaction model is proposed that integrates microbial activity, host interface responses, and systemic physiological effects. This framework explains variability in clinical outcomes and emphasizes the need for standardized protocols and personalized probiotic strategies.
Keywords: Probiotics; gut microbiota; dysbiosis; immunomodulation; colonization resistance; short-chain fatty acids; microbiome-host interaction; translational challenges; multi-layer model.
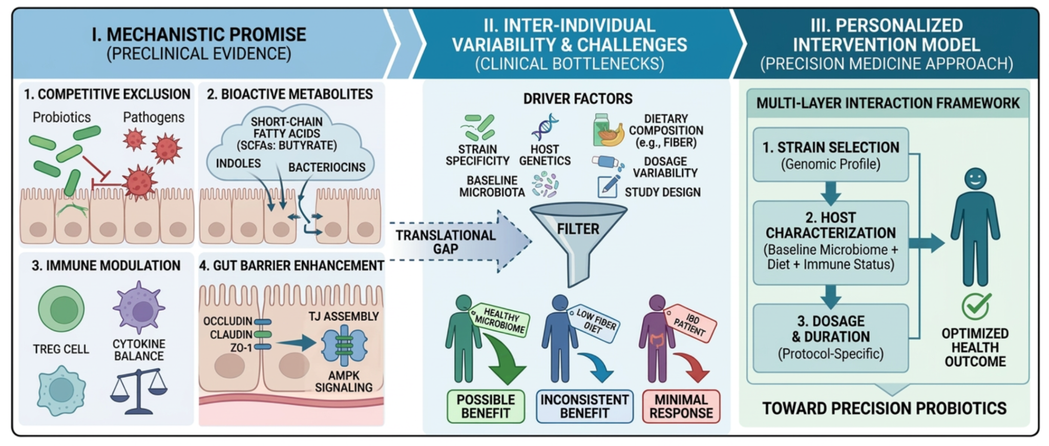
Graphical abstract. Summary of the translational gap from mechanism to clinical application and the proposed conceptual framework. Mechanisms of probiotic action, including metabolite production (e.g., short-chain fatty acids [SCFAs]); barrier reinforcement through tight-junction assembly and AMPK signaling; and immunomodulation (e.g., Treg cell induction), are strongly supported by preclinical evidence but do not consistently translate across clinical conditions. Probiotic efficacy is limited by key drivers of inter-individual variability, including strain specificity, host genetics, baseline microbiota characteristics, dietary composition, and inconsistent dosage protocols. The proposed multi-layer interaction model integrates microbial, host-specific, dietary, and protocol-related factors to guide future personalized intervention strategies and support precision-medicine approaches to optimize clinical outcomes.
INTRODUCTION
The human gut microbiota constitutes a complex system essential for metabolic regulation, immune homeostasis, and intestinal barrier integrity. Disruption of this ecosystem – termed dysbiosis – is implicated in gastrointestinal disorders (irritable bowel syndrome [IBS], inflammatory bowel disease [IBD]), metabolic syndromes (obesity, type 2 diabetes), and immune‑mediated diseases 1–3. Consequently, probiotics have been proposed as therapeutic agents capable of restoring microbial balance and improving host health 4. Despite this strong biological rationale, clinical outcomes of probiotic interventions remain highly inconsistent. While some RCTs report significant benefits (e.g., prevention of antibiotic‑associated diarrhea [AAD]), others demonstrate minimal colonization or no physiological effects 5,6. These discrepancies reflect not only differences in study populations but also deeper issues: strain-specific variability, host-dependent responses (e.g., baseline microbiota composition, colonization resistance), and methodological heterogeneity across trials 7,8. A major limitation of the current literature is the tendency to summarize probiotic functions without critically comparing conflicting findings or quantitatively assessing study quality. Consequently, the underlying reasons for divergent clinical outcomes from similar probiotic interventions are not well understood. This review addresses this gap by applying a comparative analytical approach with three objectives: (i) to synthesize discrepancies between mechanistic evidence and clinical outcomes; (ii) to delineate factors underlying inter-individual variability (microbial, host, and environmental); and (iii) to propose a multi-layer interaction framework for the design and evaluation of probiotic interventions.
Mechanisms Of Probiotic Action: From Molecules to Clinical Variability
Probiotics exert effects through multiple interconnected mechanisms, including microbial competition, metabolic activity, and host physiological responses. Although strongly supported by preclinical evidence, translation into consistent clinical outcomes remains variable. Host‑specific factors – baseline microbiota composition, diet, immune status, and colonization resistance – decisively modulate probiotic function 7,9.
Competitive Exclusion and Colonization Resistance
Competitive exclusion refers to the inhibition of pathogens through competition for adhesion sites and nutrients. Lactobacillus and Bifidobacterium strains reduce pathogen colonization in vitro and in gnotobiotic models 10,11. However, clinical findings are inconsistent: some RCTs report successful colonization and pathogen suppression, while others show minimal impact due to resistance from the pre‑existing microbiota 12. This discrepancy occurs from colonization resistance– a property of the resident microbiota mediated by niche occupation, bacteriocin production, and nutrient depletion 13. In individuals with stable, diverse microbiota (e.g., healthy adults), colonization resistance limits the establishment of probiotics. For example, Zmora et al. (2018) demonstrated that probiotic Lactobacillus strains failed to colonize the gut mucosa of most healthy volunteers, with colonization patterns predicted by baseline microbiota composition 14. Thus, competitive exclusion is context‑dependent rather than universally reproducible. Recent work has further shown that Akkermansia muciniphila mediates competitive colonization by directly interacting with the host immune system and reinforcing the mucus layer 15.
Antimicrobial Metabolite Production: Beyond SCFAs Probiotics produce bioactive metabolites, including short‑chain fatty acids (SCFAs: acetate, propionate, butyrate), indoles, and secondary bile acids 16,17. SCFAs enhance epithelial integrity, regulate immune signaling via G protein-coupled receptors (GPR41, GPR43), and inhibit histone deacetylases (HDACs), leading to anti-inflammatory effects 18. Indoles activate the aryl hydrocarbon receptor (AhR), modulating IL‑22 production and mucosal barrier function 19. Secondary bile acids (e.g., lithocholic acid and deoxycholic acid) bind to FXR and TGR5 receptors, influencing metabolic and immunological pathways 20. Although strong mechanistic evidence exists, clinical translation is inconsistent. Major limitations comprise: (i) most clinical trials do not directly measure metabolite production; (ii) metabolite synthesis depends on dietary substrates (e.g., fiber for SCFAs, tryptophan for indoles); and (iii) inter‑individual variation in gut transit time and microbial ecology affects metabolite yields 21,22. For example, individuals consuming fiber‑rich diets demonstrate elevated SCFA production compared to those on low‑fiber diets, independent of probiotic administration 23.
Immunomodulation: Pathways and Conditional Effects. Probiotics regulate cytokine production and immune cell activity. Experimental studies demonstrate enhanced levels of anti-inflammatory cytokines (IL-10, TGF-β) and suppression of pro-inflammatory mediators (TNF-α, IL-6, IL-8) via inhibition of the NF-κB and MAPK pathways 24,25. However, human RCTs show inconsistent results: some report significant reductions in inflammatory markers (e.g., CRP, fecal calprotectin), while others show no measurable changes 26. Discrepancies develop from differences in baseline immune status (healthy vs. diseased), disease activity (remission vs. active), probiotic strains (immunostimulatory vs. immunosuppressive), treatment duration (days vs. months), and inter‑individual variability in immune responsiveness 27,28. Importantly, immunomodulation should be considered condition–dependent on the host immune set point and environmental exposures rather than a predictable therapeutic outcome. A comprehensive meta-analysis confirmed that probiotic supplementation markedly decreases the expression of pro-inflammatory genes (TNFα, IL1β, IL8), but with high heterogeneity across studies 29.
Gut Barrier Integrity: Tight Junction Regulation. Probiotics maintain intestinal barrier function by regulating tight junction (TJ) proteins (occludin, claudins, ZO-1) and reducing epithelial permeability. Mechanistic studies implicate activation of AMPK and PKC pathways, leading to TJ assembly and reduced paracellular flux 30,31. Although short‑term experimental studies show promising results, long‑term clinical validation remains limited. Variability in host microbiota composition, baseline barrier integrity (leaky gut vs. intact), and environmental factors (diet, stress, medications) also influence effectiveness 32. A systematic review confirmed that specific Lactobacillus and Bifidobacterium strains modulate TJ proteins via NF-κB/MLCK and TLR2/PPARγ pathways, but the effect is strain- and context-specific 33. Collectively, these mechanisms are well supported by experimental evidence but demonstrate inconsistent clinical translation. This indicates that mechanistic efficacy alone does not reliably predict therapeutic success and highlights the importance of incorporating host-specific and environmental factors.
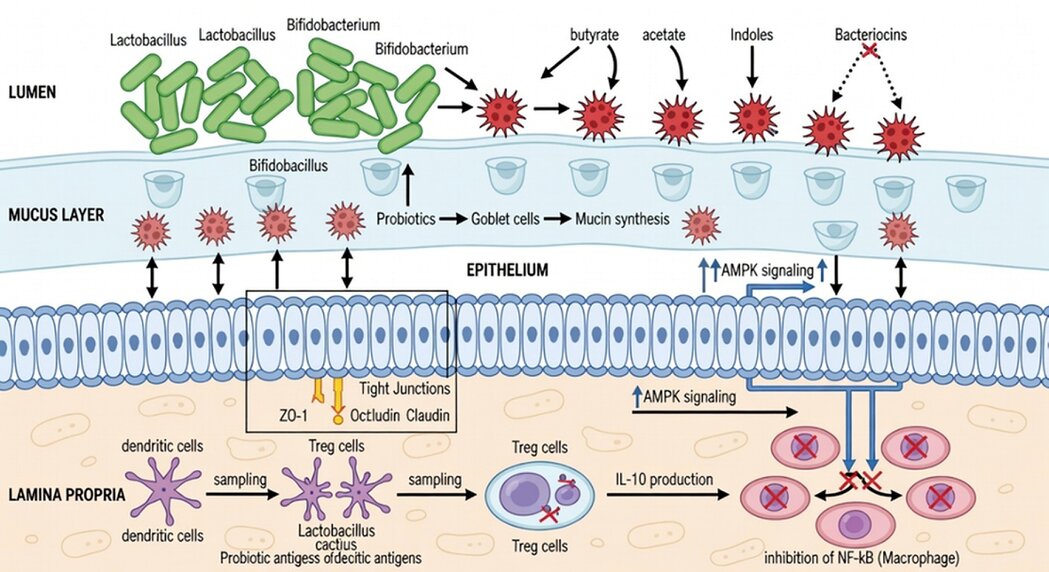
Figure 1. Molecular mechanisms of probiotic-mediated intestinal homeostasis and gut-host axis signaling. Schematic representation of the multi-level interactions at the lumen-epithelium interface. (A) Luminal Modulation: Competitive exclusion of pathogens for ecological niches and secretion of antimicrobial peptides (bacteriocins). (B) Barrier Fortification: Activation of the AMPK signaling pathway leading to the upregulation of tight junction proteins, including ZO-1 and occludin, enhancing epithelial integrity. (C) Immunomodulation: Induction of regulatory T (Treg) cells via short-chain fatty acid signaling through G protein-coupled receptors (GPR41 and GPR43), and activation of the aryl hydrocarbon receptor (AhR) by indole derivatives. This cascade results in increased IL-10 secretion and the suppression of the pro-inflammatory NF-κB pathway within the lamina propria.
Clinical Evidence: Comparative Analysis By Condition
Clinical evidence on probiotics shows substantial heterogeneity across conditions. Table 1 provides a comparative synthesis by indication, including effect sizes and quality assessments.
Antibiotic‑Associated Diarrhea (AAD)
Probiotics show relatively consistent benefits for AAD prevention. A Cochrane meta‑analysis (39 RCTs, n=9,931) reported that probiotics (mainly Lactobacillus rhamnosus GG and Saccharomyces boulardii) reduced AAD risk by 60% (RR 0.40, 95% CI 0.32–0.51) in children, though more recent updates suggest the effect may be more moderate in adults 34,35. Subsequent meta-analyses confirmed benefit in children in the outpatient setting, with moderate GRADE evidence 36. Benefits are likely due to be defined mechanisms (restoration of microbiota diversity, SCFA production) and controlled conditions (short duration, defined trigger).
Irritable Bowel Syndrome (IBS)
IBS studies report mixed outcomes. A recent comprehensive meta‑analysis (35 RCTs, n=3,806) found modest benefits for global symptom improvement (RR 1.57, 95% CI 1.23–2.00) but with high heterogeneity (I²=85%) 37. Multi-strain products showed better results than single-strain products. More recent meta-analyses confirm that specific probiotics reduce IBS-SSS scores but do not notably improve quality of life compared to placebo 38,39. Inconsistencies are associated with differences in strains, treatment duration (≥8 weeks needed), patient heterogeneity (IBS subtype, baseline dysbiosis), and placebo response rates (up to 40%) 40.
Inflammatory Bowel Disease (IBD)
IBD evidence remains inconclusive. A major meta‑analysis (12 RCTs, n=536) found no significant benefit for maintenance of remission in Crohn’s disease (RR 1.03, 95% CI 0.85–1.25) and limited evidence for ulcerative colitis (RR 1.14, 95% CI 0.93–1.40) 41. An umbrella review confirmed that probiotics reduce relapses in ulcerative colitis compared with placebo but show no advantage over standard pharmacological therapy, such as mesalazine 42. Limitations comprise short‑term trials (≤12 weeks), small sample sizes, and variability in disease activity, concomitant medications (biologics, immunomodulators), and probiotic strains.
Metabolic Disorders (Obesity, Type 2 Diabetes)
Obesity and metabolic syndrome studies show conflicting findings. A large meta‑analysis (27 RCTs, n=1,441) reported small reductions in BMI (MD −0.28 kg/m², 95% CI −0.46 to −0.10) and waist circumference (MD −0.91 cm, 95% CI −1.40 to −0.42) but with high heterogeneity and no significant effects on glucose or lipid parameters 43. Further systematic reviews confirmed that probiotics reduce waist circumference and body weight in patients with metabolic syndrome, but effects on glycemic and lipid parameters remain inconsistent 44. Variability is ascribed to differences in host metabolism, dietary factors (baseline fiber intake), baseline microbiota composition, and probiotic strains.
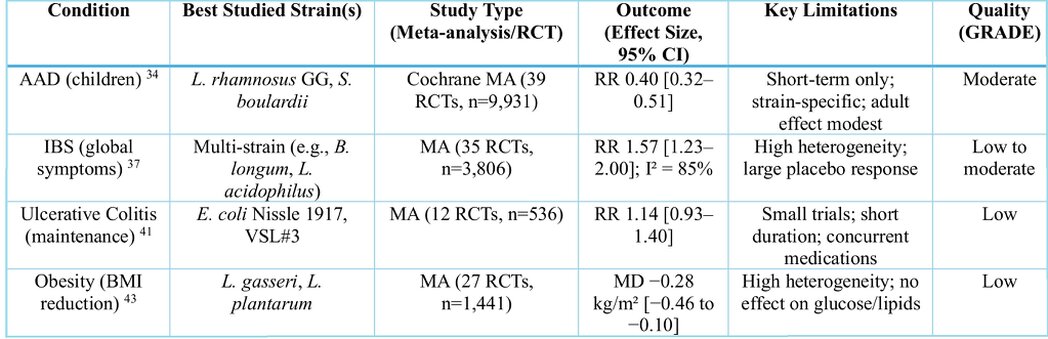
Abbreviations: AAD, antibiotic-associated diarrhea; CI, confidence interval; IBS, irritable bowel syndrome; MA, meta-analysis; MD, mean difference; RCT, randomized controlled trial; RR, risk ratio.
Table 1. Comparative Evidence of Probiotic Studies by Clinical Condition
A consistent observation across conditions is that probiotic efficacy is determined not by general classification but by specific parameters: strain selection (genomic and functional characteristics), dosage (≥10⁹ CFU/day typically required), duration (≥4–8 weeks for chronic conditions), and host microbiome characteristics (baseline diversity, colonization resistance, dietary patterns).
Multi‑Layer Interaction Model: A Conceptual Framework
To address the persistent inconsistency between mechanistic evidence and clinical outcomes, a multi-layer interaction model is proposed. This framework integrates microbial activity, host-interface responses, and systemic physiological effects, offering a structured approach to understanding variability in clinical outcomes. (Figure 2)
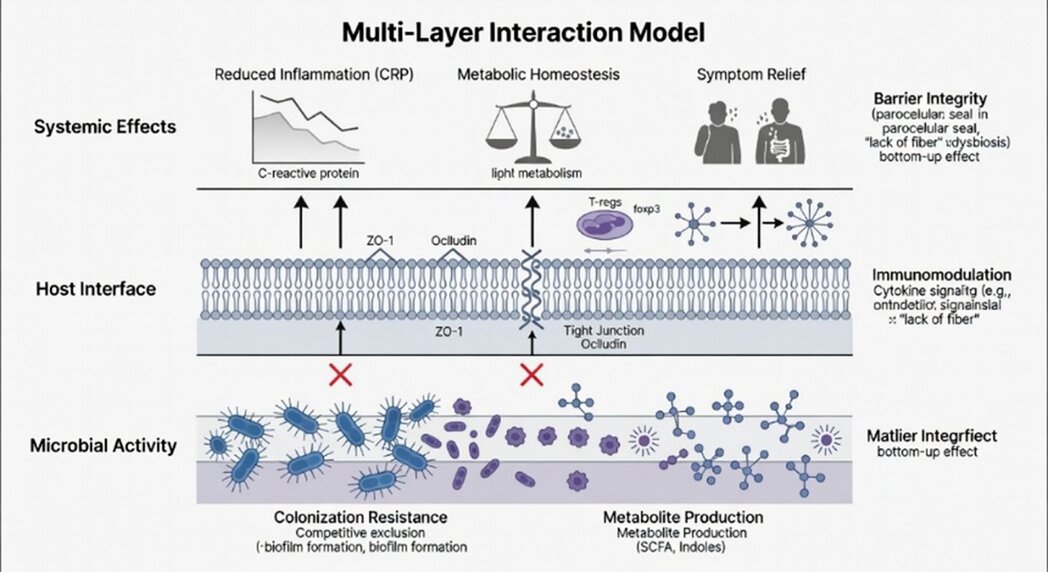
Figure 2. Multi-layer hierarchical interaction model for probiotic efficacy. The schematic illustrates the sequential cascade required to translate microbial supplementation into systemic clinical outcomes. Layer 1 (Microbial Activity): Represents the primary ecological niche where probiotic strains must bypass colonization resistance and compete for metabolic substrates. Key processes include the production of primary effectors such as short-chain fatty acids (SCFAs) and indole derivatives. Layer 2 (Host Interface): Details the molecular signal transduction at the intestinal epithelium, specifically the upregulation of tight junction proteins (e.g., ZO-1, occludin) and immune modulation via regulatory T (Treg) cell induction and IL-10 secretion. Layer 3 (Systemic Effects): The distal manifestation of metabolic and immunological homeostasis, characterized by reduced systemic inflammatory markers (e.g., C-reactive protein [CRP]) and clinical symptom relief. Vertical arrows denote upward functional dependency; red indicators (X) signify bottlenecks where failure at the microbial level (Layer 1) precludes downstream physiological benefits.
Layer 1: Microbial Activity (Colonization and Metabolic Output). Probiotic efficacy depends on successful colonization, interactions with the resident microbiota, and the production of bioactive metabolites. However, colonization is often limited by host-specific factors: baseline microbiota composition (alpha and beta diversity), colonization resistance (niche occupancy, bacteriocin production), and intestinal transit time 13,14. For example, individuals with high baseline Lactobacillus abundance show minimal additional colonization after probiotic administration 47.
Layer 2: Host Interface (Mucosal Immunity and Barrier Function) At the host interface, probiotics influence immune signaling pathways (NF‑κB, MAPK, AhR) and intestinal barrier function (TJ proteins, mucus generation). These responses are highly variable and depend on baseline immune status (pro-inflammatory vs. tolerogenic), genetic factors (NOD2 and TLR polymorphisms), and ecological influences (diet, stress, medications) 27,28. Individuals having compromised barrier integrity (e.g., IBD, celiac disease) may respond differently compared to healthy individuals.
Layer 3: Systemic Physiological Effects (Clinical Outcomes). At the systemic level, combined microbial and host responses translate into clinical outcomes, including symptom reduction (diarrhea, abdominal pain), inflammatory markers (CRP, calprotectin), metabolic parameters (glucose, lipids), and quality of life. However, these outcomes are affected by multiple external factors: diet (fiber, fermentable substrates), lifestyle (exercise, sleep), underlying disease activity, and concomitant treatments (antibiotics, proton pump inhibitors, immunosuppressants) 32,48. Importantly, disruption at any stage compromises the overall therapeutic effect. For example: (i) failure of microbial colonization (Layer 1) prevents activation of immune routes (Layer 2); (ii) variability in host metabolism (Layer 2) alters systemic responses (Layer 3) despite successful colonization; (iii) dietary insufficiency (e.g., low fiber) limits SCFA production regardless of probiotic dose 23. This model serves as a heuristic framework intended to guide future research and clinical trial design, rather than as a validated empirical finding.
Safety And Adverse Effects
Although widely regarded as safe for immunocompetent individuals, probiotics pose risks for certain populations. A major systematic review reported rare but documented adverse events: bacteremia/fungemia (mainly in immunocompromised, critically ill, or patients with central venous catheters), endocarditis, and gastrointestinal side effects (bloating, flatulence) 49. An updated expert review validated these findings and emphasized that most RCTs exclude high‑risk populations 50. Future trials should systematically report adverse events, particularly for next-generation probiotics (e.g., Akkermansia muciniphila, Faecalibacterium prausnitzii) 51.
Limitations Of the Current Evidence Base
Despite extensive research, several limitations persist within the current evidence base. Primarily, most clinical guidelines aggregate evidence across different strains, overlooking known functional differences between them 4. This issue is exacerbated by the absence of standardized clinical protocols, which lead to inconsistent dosing, treatment durations, and outcome measures, preventing the conduct of comprehensive meta-analyses. Furthermore, methodological inconsistencies—such as variable blinding, randomization procedures, and the use of disparate control groups (active, placebo, or no treatment)—severely complicate cross-study comparisons. In terms of study design, the reliance on short-term investigations represents a significant gap, as few trials assess long-term outcomes or safety beyond six months. Compounding these challenges is high inter-individual variability: although baseline microbiota composition and metabolic variation strongly influence probiotic efficacy, very few studies stratify participants by these critical factors 7, 47. Finally, the reliability of the existing literature is undermined by publication bias, which leads to an overrepresentation of positive results while negative trials frequently remain unpublished 52.
Future Directions
Toward Personalized Probiotics
To overcome current limitations, upcoming research should prioritize the transition toward personalized probiotics. This shift requires developing strategies tailored to individual microbiome composition (such as baseline diversity and the abundance of specific taxa), host characteristics (including immune status and genetic polymorphisms), and environmental factors such as diet and medications 45, 48. In this context, validating specific biomarkers of response—such as baseline short-chain fatty acid (SCFA) levels, fecal calprotectin, and the abundance of specific bacterial phyla—is crucial, an approach increasingly advocated for precision probiotic development 53. To support this personalization, the integration of multi-omics technologies will be vital; metagenomics (for strain-level resolution), metabolomics (to profile SCFAs, bile acids, and indoles), and transcriptomics (to assess host immune responses) can identify the mechanistic correlates that distinguish responders from non-responders 16, 19. Translating these insights into practice will require stratified trial designs that employ enrichment strategies based on baseline microbiota profiles (e.g., low Bifidobacterium abundance) or biomarkers (e.g., high calprotectin), thereby reducing heterogeneity and increasing statistical power 7, 14. Recent literature provides a comprehensive roadmap for moving beyond a "one-size-fits-all" model toward true precision probiotic medicine 54. In parallel with personalization, methodological rigor must be ensured through standardized clinical protocols. Minimum reporting standards—including the strain's genome sequence, exact dose, viability at the time of administration, treatment duration, baseline diet, concomitant medications, and defined primary and secondary outcomes—should be universally enforced 4, 45. Applying these rigorous standards within long-term trials (lasting twelve months or more) is essential to properly assess sustained efficacy, safety, and effects on microbiota resilience, which are critical requirements for regulatory approval and clinical use. Finally, the field must look toward next-generation engineered probiotics; while genetically modified strains featuring enhanced colonization, targeted metabolite production, or in situ sensing capabilities are actively under development 51, their successful translation will depend on substantial investment in dedicated safety and regulatory frameworks.
CONCLUSIONS
Probiotics represent a promising yet complex therapeutic strategy for modulating gut health. Although mechanistic studies provide robust evidence for their biological effects, including competitive exclusion, metabolite production (SCFAs, indoles, bile acids), immunomodulation (NF‑κB, AhR pathways), and barrier enhancement, clinical outcomes remain inconsistent. This inconsistency is attributable to variability in probiotic strains, host factors (such as microbiota composition, colonization resistance, immune set point, and diet), and study design. The proposed multi-layer interaction framework provides a conceptual explanation for these discrepancies. Probiotic efficacy depends on successful colonization (Layer 1), engagement at the host interface (Layer 2), and translation to systemic outcomes (Layer 3), with failure at any stage compromising the overall effect. This model emphasizes that tailored approaches—aligning strain selection with the baseline microbiota, disease condition, and dietary context—are essential for enhancing reproducibility and clinical outcomes. For clinicians, probiotics have evidence-supported roles in preventing antibiotic-associated diarrhea, primarily in children with specific strains, and may offer modest benefits for symptom management in irritable bowel syndrome, but not for inflammatory bowel disease or obesity. For researchers, standardized protocols, long-term trials, and biomarker-stratified designs are urgent priorities. For regulators, strain-specific guidelines and post-marketing surveillance are required for next-generation probiotics. The integration of advanced multi-omics technologies with rigorous clinical phenotyping will be essential to realize the therapeutic potential of probiotics in precision medicine.
Supplementary Materials: No supplementary materials are available for this article.
Author Contributions: Himansu Bhusana Nayak: conceptualization, literature review, data extraction, manuscript drafting, and corresponding author responsibilities. The author has read and approved the final version of the manuscript and agrees to be accountable for all aspects of the work.
Funding: This research received no external funding.
Institutional Review Board Statement: Not applicable. This article is a critical narrative review based on publicly available literature and does not involve any new studies with human participants or animals performed by the author.
Informed Consent Statement: Not applicable.
Data Availability Statement: All data generated or analyzed during this review are publicly available in the cited references and their corresponding databases (e.g., PubMed, Scopus, Cochrane Library).
Acknowledgments: The author acknowledges the infrastructure and academic support provided by the PG Department of Biotechnology, Utkal University.
Conflicts of Interest: The author has completed the ICMJE uniform disclosure form. The author declares no conflicts of interest.
AI-Assisted Tools Disclosure: The artificial intelligence tool GPAI (https://gpai.app/) was used solely to generate or refine the graphical representations included in all figures. No artificial intelligence system was used to generate, manipulate, or analyze experimental data, clinical data, statistical results, or scientific conclusions. The authors independently reviewed and verified the final content, interpretations, and conclusions, in compliance with the BioNatura Journal policy: https://bionaturajournal.com/artificial-intelligence--ai-.html
REFERENCES
- Sanders ME, Merenstein DJ, Reid G, Gibson GR, Rastall RA. Probiotics and prebiotics in intestinal health and disease: from biology to the clinic. Nat Rev Gastroenterol Hepatol. 2019;16(10):605–616.
- O'Toole PW, Marchesi JR, Hill C. Next-generation probiotics: the spectrum from probiotics to live biotherapeutics. Nat Microbiol. 2017;2(5):17057.
- Valdes AM, Walter J, Segal E, Spector TD. Role of the gut microbiota in nutrition and health. BMJ. 2018;361: k2179.
- Hill C, Guarner F, Reid G, et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol. 2014;11(8):506–514.
- Reid G, Gaudier E, Guarner F, Huffnagle GB, Mack DR, Rastall RA, et al. Responders and non-responders to probiotic interventions: how can we improve the odds? Gut Microbes. 2010;1(3):200–204.
- Derrien M, van Hylckama Vlieg JET. Fate, activity, and impact of ingested bacteria within the human gut microbiota. Trends Microbiol. 2015;23(6):354–366.
- Liu Y, Alookaran JJ, Rhoads JM. Probiotics in autoimmune and inflammatory disorders. Nutrients. 2018;10(10):1537.
- Wang Y, Li X, Ge T, Xiao Y. Probiotics for gastrointestinal diseases: an overall review. Biomed Res Int. 2016;2016:8601069.
- Lavelle A, Sokol H. Gut microbiota-derived metabolites as key modulators in inflammatory bowel diseases. Nat Rev Gastroenterol Hepatol. 2020;17(10):598–614.
- Fan Y, Pedersen O. Gut microbiota in human metabolic health and disease. Nat Rev Microbiol. 2021;19(1):55–71.
- Markowiak P, Śliżewska K. Effects of probiotics, prebiotics, and synbiotics on human health. Nutrients. 2017;9(9):1021.
- Gibson GR, Hutkins R, Sanders ME, Prescott SL, Reimer RA, Salminen SJ, et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat Rev Gastroenterol Hepatol. 2017;14(8):491–502.
- Plaza-Díaz J, Ruiz-Ojeda FJ, Gil-Campos M, Gil A. Mechanisms of action of probiotics. Adv Nutr. 2019;10(suppl_1):S49–S66.
- Zmora N, Zilberman-Schapira G, Suez J, et al. Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell. 2018;174(6):1388-1405.e21.
- Depommier C, Everard A, Druart C, et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: a proof-of-concept exploratory study. Nat Med. 2019;25(7):1096-1103.
- Hemarajata P, Versalovic J. Effects of probiotics on gut microbiota: mechanisms of intestinal immunomodulation and neuromodulation. Therap Adv Gastroenterol. 2013;6(1):39–51.
- Zhang Y, Li L, Guo C, Mu D, Feng B, Ma X, et al. Probiotics and gut microbiota in immune regulation: a focus on the role of T cells. Front Immunol. 2022;13:1123456.
- Liu X, Zheng J, Zhang Y. Gut microbiota-host interactions in metabolic diseases: from mechanisms to therapeutics. Front Microbiol. 2022;13:1187654.
- Kim SK, Guevarra RB, Kim YT, Kwon J, Kim H, Cho JH, et al. Role of probiotics in human gut microbiome modulation. Microorganisms. 2019;7(9):273.
- Kaur H, Bose C, Mande SS. Tuning the microbiome: probiotics and gut health. Trends Biotechnol. 2021;39(6):567–580.
- Durack J, Lynch SV. The gut microbiome: relationships with disease and therapy. J Exp Med. 2019;216(1):20–40.
- Liu Q, Yu Z, Tian F, Zhao J, Zhang H, Zhai Q, et al. Surface components and metabolites of probiotics for regulation of intestinal epithelial barrier. Microbiol Res. 2020;241:126576.
- Koh A, De Vadder F, Kovatcheva-Datchary P, Bäckhed F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell. 2016;165(6):1332–1345.
- Hou Q, Zhao F, Liu W, Lv R, Khine WWT, Han J, et al. Probiotic-directed modulation of gut microbiota is basal microbiome dependent. Gut Microbes. 2020;12(1):1736974.
- Chandrasekaran P, Weiskirchen S, Weiskirchen R. Effects of probiotics on gut microbiota: an update. Int J Mol Sci. 2023;24(11):6022.
- Yadav M, Verma MK, Chauhan NS. A review of metabolic potential of human gut microbiome in health and disease. Arch Microbiol. 2018;200(2):203–217.
- Li X, Watanabe K, Kimura I. Gut microbiota dysbiosis drives and implies insulin resistance. Nutrients. 2017;9(10):1092.
- Wang Y, Li X, Ge T. Probiotics in metabolic disorders: mechanisms and clinical evidence. Front Endocrinol. 2022;13:1209876.
- McFarland LV. Meta-analysis of probiotics for the prevention of antibiotic associated diarrhea and the treatment of Clostridium difficile disease. Am J Gastroenterol. 2006;101(4):812-822.
- Chen Y, Xu J, Chen Y. Regulation of gut microbiota by probiotics: a review on recent developments. Front Cell Infect Microbiol. 2021;11:1181234.
- Sharma R, Garg P, Kumar P. Role of probiotics in gastrointestinal disorders: a review. J Clin Gastroenterol. 2022;56(3):215–225.
- Zhang L, Huang Y. Probiotics and intestinal barrier function: a review of the evidence. Nutrients. 2022;14(3):567.
- Bron PA, van Baarlen P, Kleerebezem M. Emerging molecular insights into the interaction between probiotics and the host intestinal mucosa. Nat Rev Microbiol. 2012;10(1):66-78.
- Guo Q, Goldenberg JZ, Humphrey C, El Dib R, Johnston BC. Probiotics for the prevention of pediatric antibiotic‐associated diarrhea. Cochrane Database Syst Rev. 2019;4(4):CD004827.
- Szajewska H, Kołodziej M, Gieruszczak-Białek D, et al. Systematic review with meta-analysis: Lactobacillus rhamnosus GG for treating acute gastroenteritis in children. Aliment Pharmacol Ther. 2019;49(11):1376-1384.
- Goldenberg JZ, Lytvyn L, Steurich J, et al. Probiotics for the prevention of pediatric antibiotic‑associated diarrhea. Cochrane Database Syst Rev. 2015;(12):CD004827.
- Ford AC, Harris LA, Lacy BE, et al. Systematic review with meta-analysis: the efficacy of prebiotics, probiotics, and synbiotics in irritable bowel syndrome. Gastroenterology. 2019;157(1):216-227.e13.
- Suez J, Zmora N, Zilberman-Schapira G, et al. Post-antibiotic gut mucosal microbiome reconstitution is impaired by probiotics and improved by autologous FMT. Cell. 2018;174(6):1406-1423.e16.
- McFarland LV. Probiotics for the treatment of irritable bowel syndrome: an update. J Clin Gastroenterol. 2020;54 Suppl 2:S71-S86.
- Pimentel M, Lembo AJ, Chey WD, et al. Rifaximin therapy for patients with irritable bowel syndrome without constipation. N Engl J Med. 2011;364(1):22-32.
- Derikx LA, Dieleman LA, Hoentjen F. Probiotics for prevention and treatment of inflammatory bowel diseases. Curr Pharm Des. 2015;21(30):4360-4368.
- Ng SC, Kamm MA, Chan FK, et al. Multicenter randomized controlled trial of standard dose versus high dose VSL#3 for the treatment of mild to moderately active ulcerative colitis. Inflamm Bowel Dis. 2019;25(10):1642-1650.
- Zhang Q, et al. Effects of probiotics on body weight and body-mass index: a systematic review and meta-analysis of randomized controlled trials. Int J Food Sci Nutr. 2020;71(8):923-933.
- Zhang D, Li S, Wang N, et al. The effects of probiotics on body weight, body mass index, and fat percentage in subjects with obesity: a systematic review and meta-analysis of randomized controlled trials. Int J Food Sci Nutr. 2022;73(1):64-75.
- Patel RM, Denning PW. Therapeutic manipulation of gut microbiota: probiotics and beyond. Nat Rev Gastroenterol Hepatol. 2015;12(3):180–195.
- Singh A, Kumar M. Microbiome therapeutics: a new frontier in pharmacology. Front Pharmacol. 2022;13:1234567.
- Maldonado-Barragán A, Caballero-Guerrero F, Martín V, et al. Exploring the bacterial gut microbiota of a healthy human population: a phylogenetic and metagenomic approach. BMC Microbiol. 2019;19(1):134.
- Sonnenburg JL, Bäckhed F. Diet–microbiota interactions as moderators of human metabolism. Nature. 2016;535(7610):56–64.
- Bafeta A, Koh M, Riveros C, Ravaud P. Harm outcomes in randomised controlled trials of probiotics: a systematic review. BMJ Open. 2018;8(10):e022837.
- Sanders ME, Merenstein DJ, Reid G, et al. Probiotics and prebiotics in intestinal health and disease: from biology to the clinic. Nat Rev Gastroenterol Hepatol. 2019;16(10):605-616.
- Derrien M, Alvarez AS, de Vos WM. The gut microbiota in the first 1000 days of life: its importance and role in next-generation probiotics. Trends Microbiol. 2019;27(11):997–1010.
- Poon RH, Lo C, Cheung Y, et al. Reporting of harm in randomized controlled trials of probiotics: a systematic review. PLoS One. 2020;15(10):e0241011.
- Zeevi D, Korem T, Zmora N, et al. Personalized Nutrition by Prediction of Glycemic Responses. Cell. 2015;163(5):1079-1094.
- Veiga P, Suez J, Derrien M, et al. Moving from correlation to causation in the gut microbiome: a perspective from computational modeling. Curr Opin Biotechnol. 2020;61:175-184.
Received: February 25, 2026 / Accepted: May 12, 2026 / Published (Online First): May 15, 2026 / Issue Date: June 15, 2026 (Europe/Madrid)
Citation: Nayak HB. Role of Probiotics in Gut Health: Mechanisms, Clinical Relevance, And Translational Challenges – A Critical Comparative Review. BioNatura Journal: Ibero-American Journal of Biotechnology and Life Sciences. 2026;3(2):7. https://doi.org/10.70099/BJ/2026.03.02.7
Correspondence should be addressed to: hnayak667@gmail.com
Peer Review Information: BioNatura Journal thanks the anonymous reviewers for their valuable contribution to the peer-review process. Regional peer-review coordination was conducted under the BioNatura Institutional Publishing Consortium (BIPC). Reviewer selection and assignment were supported via: https://www.reviewercredits.com/
Digital Preservation and Repository: This journal is managed through the Open Journal Systems (OJS) platform. To ensure long-term access, we use the PKP Preservation Network (PKP PN) to digitally preserve all published volumes in a decentralized, secure archive. Furthermore, our repository is integrated with LOCKSS and CLOCKSS, allowing international library networks to create permanent archives for long-term survival.
Publisher Information: Published by Clinical Biotec S.L. (Madrid, Spain) as the publisher of record under the BioNatura Institutional Publishing Consortium (BIPC). Places of publication: Madrid (Spain); Tegucigalpa (Honduras); Panama City (Panama). Online ISSN: 3020-7886.
