Microbiological characterization of raw goat milk from a small municipality in Brazil: identification of Gram-positive cocci by MALDI-TOF MS
Gustavo Luis de Paiva Anciens Ramos 



Janaína dos Santos Nascimento
Janaína dos Santos Nascimento
1 Universidade Federal Fluminense; gustavoanciens@id.uff.br.
2 Instituto Federal de Educação, Ciência e Tecnologia do Rio de Janeiro; emily.silva.etm2020.1@gmail.com; lavinia.araujo.etm2021.1@gmail.com,soares.marcelo2020@gmail.com, janaina.nascimento@ifrj.edu.br.
* Correspondence: gustavoanciens@id.uff.br
ABSTRACT
Goat milk is a high-nutritional-value food, but it can also harbor several pathogenic microorganisms. It is not uncommon for this food to be consumed raw, which can pose health risks to consumers. In this study, we sought to verify the diversity of Gram-positive cocci (GPC), especially Staphylococcus sp., in raw milk samples obtained in the region of Itaocara, RJ. Microbial identification of the nine samples was performed by mass spectrometry (MALDI-TOF) after isolation in selective culture media. MALDI-TOF analysis of the isolates from raw goat milk samples revealed the predominance of Staphylococcus spp., with emphasis on S. aureus and S. caprae. GPCs of other genera were also detected. Surprisingly, some of these microorganisms were isolated on selective media for Gram-negative bacteria, highlighting the importance of accurate confirmatory identification methods such as MALDI-TOF. The detection of S. aureus, a relevant foodborne pathogen, and other species with spoilage or pathogenic potential, such as S. caprae and Mammaliicoccus sciuri, in six of the nine samples highlights potential microbiological risks associated with locally consumed raw goat milk. On the other hand, the presence of lactic acid bacteria (Lactococcus lactis, L. garvieae, and Enterococcus sp.), with technological potential for producing dairy products, was also observed. The recovery of Gram-positive cocci from media selective for Gram-negative bacteria reinforces the limitations of culture-based selectivity and highlights the importance of confirmatory identification methods such as MALDI-TOF MS.
Keywords: Staphylococcus aureus, dairy product, mass spectrometry
Keywords: Staphylococcus aureus, dairy product, mass spectrometry
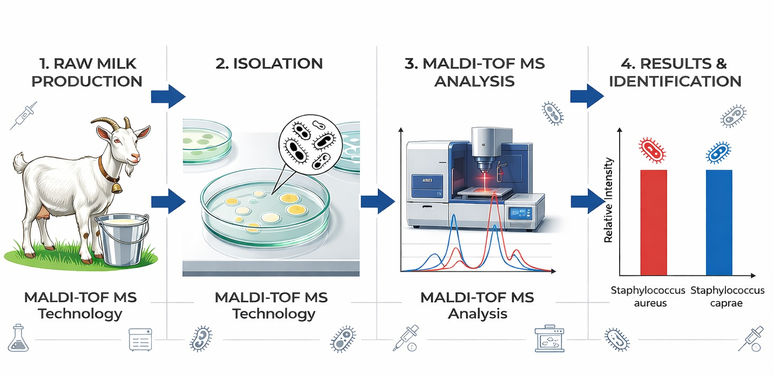
Graphical Abstract. Comprehensive microbiological workflow for the characterization of raw goat milk. The study integrated: (1) Sampling from small-scale production in Itaocara, RJ; (2) Microbial isolation using selective media; (3) Advanced identification via MALDI-TOF MS technology; and (4) Accurate species-level results highlighting the prevalence of Staphylococcus aureus and S. caprae. Image generated using AI-assisted tools (GPAI).
INTRODUCTION
Goat farming has been growing, driven by the demand for more natural products and differentiated dairy foods 1. In this context, goat's milk stands out for its unique nutritional profile, including high levels of vitamins, minerals, and medium-chain fatty acids, making it a food appreciated worldwide. Additionally, it is an option for individuals with lactose intolerance or cow's milk allergy 2.
However, the marketing and consumption of raw goat's milk, a common practice in some regions, raises significant concerns regarding microbiological safety. Raw milk, as it does not undergo heat pasteurization, can harbor a diverse microbiota, including Gram-positive bacteria with pathogenic and deteriorative potential 3. Among these bacteria, the Staphylococcus genus has been frequently isolated from raw milk of different species, posing a risk to public health due to the production of heat-stable enterotoxins (responsible for food poisoning), biofilm formation, and antimicrobial resistance 4,5. The presence of other Gram-positive cocci can also cause disease in humans, goats, and other domestic ruminants (clinical and subclinical mastitis), as well as compromising the sensory quality and shelf life of milk and its derivatives 6-8.
On the other hand, other GPC, such as Lactococcus and Enterococcus, play a crucial role in lactic fermentation, contributing to the development of characteristic flavor, aroma, and texture in cheeses and other fermented products 9. Therefore, understanding the microbial diversity in raw goat's milk is essential for assessing health risks and identifying the biotechnological potential for producing high-quality artisanal foods.
In this context, the present study aimed to analyze the presence and diversity of potentially pathogenic or spoilage GPC in raw goat's milk samples sold by small producers in the Itaocara region, in the northwest of the state of Rio de Janeiro, and consumed by the local population.
MATERIAL AND METHODS
Sampling
Nine samples of raw goat's milk (Table 1), commonly sold and consumed in natura by the local population, were obtained from small producers in the municipality of Itaocara, in the northwest of the State of Rio de Janeiro, between July and September 2024. The samples come from family farms, which are a major activity in the community of about 23,000 people. The samples were transported under refrigeration to the IFRJ Microbiology Laboratory.
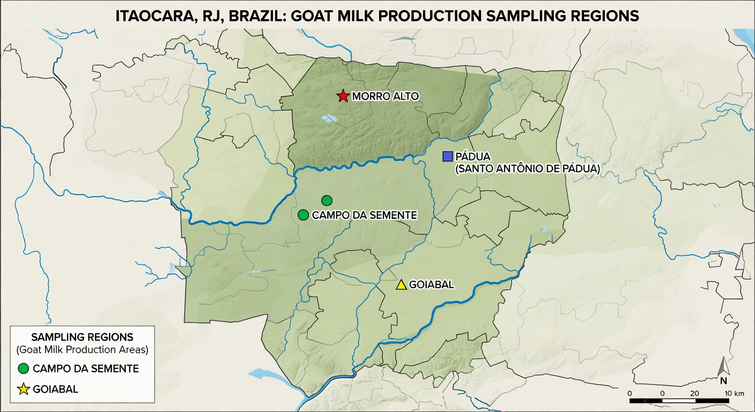
Figure 1. Distribution of microbial species identified in raw goat milk samples from Itaocara, RJ. Staphylococcus aureus was the most prevalent species (36.1%), followed by Staphylococcus caprae (16.6%).

Table 1. Samples of raw goat milk used in this work
Obtaining the isolates
To obtain GPC, primarily to isolate Staphylococcus sp., the methodology described in ISO 6888-1: 1999 10 was used. It is known, however, that other GPCs, generally related to Staphylococcus sp., can also be isolated using this methodology. Samples were diluted to 10-3, plated on Baird-Parker agar (Himedia, São Paulo, Brazil), supplemented with egg yolk emulsion and potassium tellurite, and incubated at 37°C for 48 h. Typical and atypical colonies (suspected coagulase-positive and coagulase-negative S.) were selected and subcultured on a non-selective medium (Casoy agar, Himedia, São Paulo, Brazil). The plates were incubated at 37°C for 18 to 24h. For a parallel study, the same samples were also inoculated on selective and differential culture media for Gram-negative bacteria (Leeds Agar, Cetrimide Agar, and Tryptone Bile X-glucuronide Agar).
Identification of isolates
The identification of the isolates was performed by MALDI-TOF (Matrix-Assisted Laser Desorption/Ionization – Time of Flight) mass spectrometry on a mass spectrometer (Microflex LT, Bruker, United States) at the Medical Microbiology Research Laboratory of the Paulo de Góes Institute of Microbiology, at the Federal University of Rio de Janeiro (UFRJ). Bacterial cultures previously grown on Casoy agar were inoculated in duplicate onto the equipment's metal plate, followed by the addition of 1 μl of 70% formic acid for cell lysis. After the acid dried, the same volume of a matrix of α-cyano-4-hydroxycinnamic acid was applied to crystallize the sample at the end of the process. The plate was then inserted into the equipment, which had been previously calibrated with a control strain of Escherichia coli, and the Biotype 3.1 software generated an identification for each isolate. Isolates with species-level accuracy were identified when their spectral similarity score exceeded 2.0.
RESULTS AND DISCUSSION
From the colonies obtained from the microbiological analysis of nine samples of raw goat's milk from different small producers in the region of Itaocara, RJ, some isolates were selected and subjected to MALDI-TOF mass spectrometry analysis. Those that did not show growth in the non-selective medium or did not present precise identification of genus or species were discarded. The isolates were identified as Staphylococcus aureus (n=13; 36.1%), S.caprae (n=6; 16.6%), Staphylococcus sp. (n=4; 11.1%), Enterococcus faecium (n=3; 8.3%), Macrococcus caseolyticus (n=3; 8.3%), Mammaliicoccus sciuri (n=2; 5.6%), S. kloosii (n=1; 2.8%), Enterococcus sp. (n=1; 2.8%), Rothia koreensis (n=1; 2.8%), Lactococcus lactis (n=1; 2.8%) and L. garvieae (n=1; 2.8%). Percentages are presented as descriptive values of the isolates analyzed in this study. The identification of the isolates from the goat milk sample is presented in Table 2.
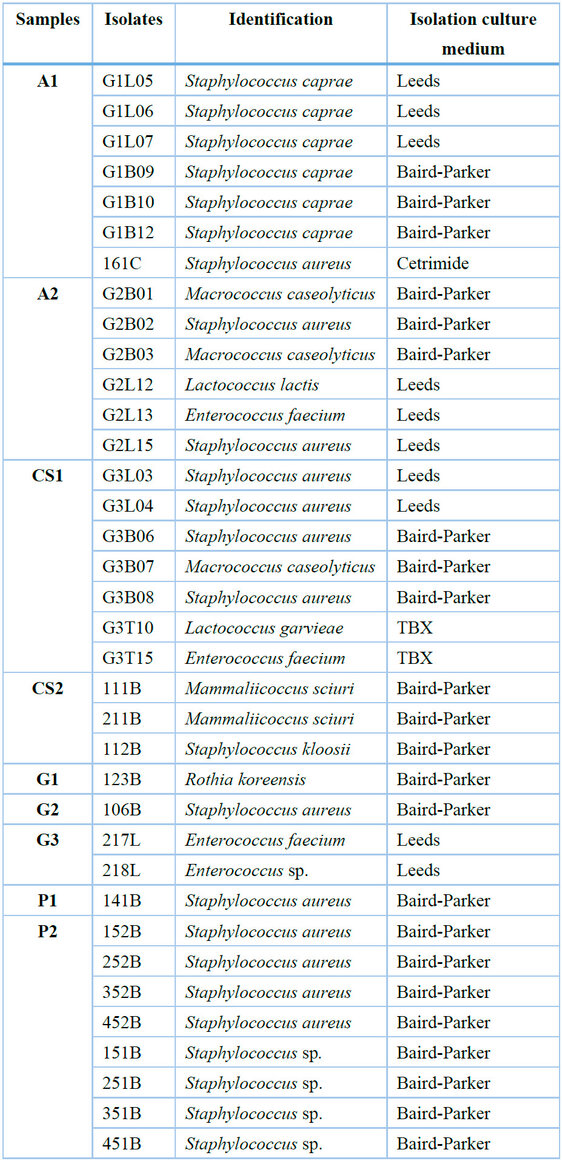
Table 2. Gram-positive cocci isolated from the raw goat milk samples. The isolate codes refer to morphological information about the colonies used to organize the study.
Figure 2. Distribution of microbial species identified in raw goat milk samples from Itaocara, RJ. Staphylococcus aureus was the most prevalent species (36.1%), followed by Staphylococcus caprae (16.6%).
Some isolates were recovered not only on Baird-Parker agar but also on selective and differential media intended for Gram-negative bacteria (Leeds, Cetrimide, and TBX agars), originally used for a parallel study.
The recovery of Gram-positive cocci on these media represents a relevant methodological finding. Although selective media are designed to suppress non-target microorganisms, their specificity is not absolute 11. In this study, the growth of Staphylococcus, Enterococcus, and Lactococcus species on media intended for Gram-negative bacteria demonstrates the practical limitations of phenotypic selectivity. These results indicate that relying solely on culture-based characteristics may lead to misinterpretation of microbial identity.
In this context, MALDI-TOF mass spectrometry proved essential for the accurate identification of isolates recovered under these atypical culture conditions, as it supports diagnostic and research microbiology by enabling the reliable identification of microorganisms based on their unique protein profiles. This technique overcomes the limitations of conventional phenotypic methods, which can be time-consuming and, in certain situations, not discriminatory enough to distinguish phylogenetically close species, mainly when growth occurs in atypical culture media 12. For example, MALDI-TOF identification of Lactococcus garvieae, Enterococcus faecium, and the genera Macrococcus, Mammaliicoccus, and Rothia provides reliable information on the presence of these species in the samples, regardless of the initial isolation medium. It is worth noting that isolates that did not yield conclusive identification are expected and emphasize the importance of continuously updating the MALDI-TOF reference databases 13, especially for non-clinical samples.
The predominance of Staphylococcus isolates (63.8%) was expected; however, the diversity of other isolated species demonstrates the complexity of the microbiota of raw goat milk and underscores the importance of more in-depth studies to characterize these microorganisms functionally. The most prevalent species among the isolates, S. aureus, was detected in six of the nine samples analyzed. This finding is consistent with previous studies that reported the frequent occurrence of S. aureus in raw milk from different animal species 8. The presence of this microorganism in raw milk is considered a significant public health risk, since this bacterium is recognized as an important foodborne pathogen capable of producing heat-stable enterotoxins that can cause food poisoning even after milk heat treatment 14, and the spread of antibiotic-resistant S. aureus strains further aggravates this concern 15. However, in the present study, no enterotoxin production or virulence factors were evaluated.
In contrast, S. caprae was isolated in only one sample (A1). Although generally considered to have low pathogenic potential, some strains of coagulase-negative staphylococci, such as S. caprae, are more associated with subclinical mastitis than with clinical mastitis, which is commonly associated with S. aureus 16. Furthermore, S. caprae can occasionally cause human infections 17. The presence of other Staphylococcus species underscores the importance of monitoring the microbiological quality of raw goat's milk, as some species of this genus may exhibit pathogenic and/or deteriorative characteristics.
Macrococcus caseolyticus, isolated from samples A2 and CS1, is known for its association with changes in the flavor of milk and dairy products that can generate undesirable flavors. However, regarding the safety of using macrococcal strains in food, two important criteria should be evaluated: the presence of antimicrobial resistance determinants and their transfer to other bacteria, since this genus is closely related to Staphylococcus sp. 7, 18. The presence of Mammaliicoccus sciuri, previously known as S. sciuri, in sample CS2 may indicate potential failures in herd hygiene and health practices during production, since this species is frequently found on animals' skin and may contaminate milk. Its importance lies in its being a bacterium generally resistant to antibiotics, capable of colonizing animals and humans, and considered a potentially opportunistic pathogen 19, 20.
Rothia koreensis, found in sample G1, is a bacterium that has not yet been studied; until 2018, it was still classified in the genus Kocuria. The few studies that cite the genus Rothia associate it with infections in immunocompromised or immunocompetent individuals 21, but its role in dairy foods is limited to its former nomenclature, Kocuria koreensis, with few records of its isolation from raw bovine milk and cheeses 22, 23.
Lactic acid bacteria (LAB) were also observed. These microorganisms, commonly found in milk, are mostly non-pathogenic and have great potential for technological application in the production of dairy products. Lactococcus lactis and L. garvieae, species widely used as starter cultures in cheese production, were isolated in samples A2 and CS1, respectively. These microorganisms play a key role in milk acidification and the development of desirable aromatic compounds 24. The detection of Enterococcus faecium in three samples (A2, CS1, and G3) is also relevant, as this species can contribute to the ripening and flavor of dairy products such as cheese 25. The isolation of these LAB on Gram-negative media (Leeds agar and TBX agar) further highlights the importance of accurate identification techniques such as MALDI-TOF.
This study has some limitations that should be acknowledged. The number of samples analyzed was relatively small, and sampling was restricted to specific small towns, limiting the generalizability of the findings. In addition, this work has a descriptive character, focusing on the isolation and identification of microorganisms. No analyses of virulence genes, enterotoxin production, or antimicrobial resistance profiles were performed. Furthermore, the technological properties of the isolates were not experimentally evaluated. It should also be considered that the culture conditions and the use of selective media may have influenced the spectrum of microorganisms recovered. Therefore, any discussion regarding pathogenic potential, resistance traits, or technological applications is based on previous literature and should not be interpreted as characteristics demonstrated in the isolates analyzed in this study.
Future studies should expand the sampling scope to include a larger number of samples and broader geographic regions, enabling a more comprehensive assessment of the microbiological quality of raw goat milk. Quantitative microbiological analyses would also be valuable for better characterizing contamination levels. In addition, molecular approaches targeting virulence factors and enterotoxin genes, as well as antimicrobial resistance profiling, would provide further insight into the potential risks associated with the identified microorganisms. Finally, evaluating the technological properties of selected lactic acid bacteria could help explore their potential applications in dairy production.
CONCLUSIONS
This study provides a descriptive overview of Gram-positive cocci present in raw goat milk from small-scale producers in a specific region of Brazil, revealing a diverse microbiota dominated by Staphylococcus species. The detection of S. aureus in several samples indicates potential microbiological risks at the local level, although broader generalizations cannot be inferred from the limited sample size.
A key finding of this study was the recovery of Gram-positive cocci from culture media intended for Gram-negative bacteria, highlighting the limitations of phenotypic selectivity in routine microbiological analysis. This result reinforces the need for confirmatory identification techniques. In this context, MALDI-TOF MS proved to be a robust and reliable tool for accurate microbial identification, particularly when isolates are obtained under atypical or non-selective growth conditions.
Author Contributions: Gustavo L. P. A. Ramos: Data analysis and interpretation, article writing, and critical review of intellectual content. Emily C. C. Silva, Lavínia F. Araújo, and Marcelo S. Moraes: Experiment execution, data analysis, and interpretation. Janaína S. Nascimento: Concept and design, data analysis and interpretation, article writing, and critical review of intellectual content. All authors approve the final version to be published and agree to be responsible for all aspects of the work.
Funding: This study was supported by Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq).
Institutional Review Board Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study does not involve human participants, and therefore, Institutional Review Board approval and informed consent were not required for its execution.
Informed Consent Statement: Not applicable. This study does not involve human subjects.
Data Availability Statement: The data supporting the findings of this study will be made available by reasonable request to the corresponding author.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form. The authors have no conflicts of interest to declare.
AI-Assisted Tools Disclosure: The artificial intelligence tool GPAI (https://gpai.app/) was used solely to generate the images included in this study. No artificial intelligence system was used to generate, manipulate, or analyze experimental data or statistical results. The authors independently verified all results, analyses, and conclusions, in compliance with the BioNatura Journal policy: https://bionaturajournal.com/artificial-intelligence--ai-.html
REFERENCES
1. Moraes MS, Ramos GLP, Nascimento JS (2025) Benefits of goat, sheep and buffalo milk: A concise review. Int J Vet Sci Anim Husb 10:150–155.
2. Roy D, Ye A, Moughan PJ, Singh H (2020) Composition, structure, and digestive dynamics of milk from different species—A review. Front Nutr 7:577759. https://doi.org/10.3389/fnut.2020.577759
3. Fusco V, Chieffi D, Fanelli F, Logrieco AF, Cho GS, Kabisch J, Arioli F, Papademas P, De Angelis M, Franz CM (2020) Microbial quality and safety of milk and milk products in the 21st century. Compr Rev Food Sci Food Saf 19:2013–2049. https://doi.org/10.1111/1541-4337.12568
4. Ruiz P, Barragán I, Seseña S, Palop ML (2016) Is staphylococci population from milk of healthy goats safe? Int J Food Microbiol 238:146–152. https://doi.org/10.1016/j.ijfoodmicro.2016.09.006
5. Wiśniewski P, Gajewska J, Zadernowska A, Chajęcka-Wierzchowska W (2024) Identification of the enterotoxigenic potential of Staphylococcus spp. from raw milk and raw milk cheeses. Toxins 16:17. https://doi.org/10.3390/toxins16010017
6. Machado MAA, Ribeiro WA, Toledo VS, Ramos GLPA, Vigoder HC, Nascimento JS (2020) Antibiotic resistance and biofilm production in Gram-positive catalase-positive cocci isolated from Brazilian pasteurized milk. J Food Qual Hazards Control 7:67–74.
7. Ramos GLPA, Vigoder HC, Nascimento JS (2021) Technological applications of Macrococcus caseolyticus and its impact on food safety. Curr Microbiol 78:11–16. https://doi.org/10.1007/s00284-020-02264-5
8. Ruiz-Romero RA, Vargas-Bello-Pérez E (2023) Non-aureus staphylococci and mammaliicocci as a cause of mastitis in domestic ruminants: A systematic review. Vet Res Commun 47:1067–1084. https://doi.org/10.1007/s11259-023-10135-0
9. Terzić-Vidojević A, Veljović K, Popović N, Tolinački M, Golić N (2021) Enterococci from raw-milk cheeses: Current knowledge on safety, technological, and probiotic concerns. Foods 10:2753. https://doi.org/10.3390/foods10112753
10. International Organization for Standardization (1999) ISO 6888-1:1999. Microbiology of food and animal feeding stuffs—Horizontal method for the enumeration of coagulase-positive staphylococci (Staphylococcus aureus and other species)—Part 1: Technique using Baird-Parker agar medium.
11. Bonnet M, Lagier JC, Raoult D, Khelaifia S (2019) Bacterial culture through selective and non-selective conditions: the evolution of culture media in clinical microbiology. New Microbes New Infect 34:100622. https://doi.org/10.1016/j.nmni.2019.100622
12. Calderaro A, Chezzi C (2024) MALDI-TOF MS: A reliable tool in the real life of the clinical microbiology laboratory. Microorganisms 12:322. https://doi.org/10.3390/microorganisms12020322
13. Rychert J (2019) Benefits and limitations of MALDI-TOF mass spectrometry for the identification of microorganisms. J Infectiol Epidemiol 2:1510109. https://doi.org/10.23937/2474-3658/1510109
14. Grispoldi L, Karama M, Armani A, Hadjicharalambous C, Cenci-Goga BT (2021) Staphylococcus aureus enterotoxin in food of animal origin and staphylococcal food poisoning risk assessment from farm to table. Ital J Anim Sci 20:677–690. https://doi.org/10.1080/1828051X.2021.1901263
15. Zhang J, Wang J, Jin J, Li X, Zhang H, Shi X, Zhao C (2022) Prevalence, antibiotic resistance, and enterotoxin genes of Staphylococcus aureus isolated from milk and dairy products worldwide: A systematic review and meta-analysis. Food Res Int 162:111969. https://doi.org/10.1016/j.foodres.2022.111969
16. Nelli A, Voidarou C, Venardou B, Fotou K, Tsinas A, Bonos E, Fthenakis GC, Skoufos I, Tzora A (2022) Antimicrobial and methicillin resistance pattern of potential mastitis-inducing Staphylococcus aureus and coagulase-negative staphylococci isolates from dairy goats. Biology 11:1591. https://doi.org/10.3390/biology11111591
17. Mastroianni A, Greco S, Vangeli V, Mauro MV, Manfredi R (2023) Invasive diseases due to fatal Staphylococcus caprae infection: A case series and literature review. J Infect Dis Epidemiol 9:298. https://doi.org/10.23937/2474-3658/1510298
18. Mazhar S, Kilcawley KN, Hill C, McAuliffe O (2020) A systems-wide analysis of proteolytic and lipolytic pathways uncovers the flavor-forming potential of the Gram-positive bacterium Macrococcus caseolyticus subsp. caseolyticus. Front Microbiol 11:1533. https://doi.org/10.3389/fmicb.2020.01533
19. Madhaiyan M, Wirth JS, Saravanan VS (2020) Phylogenomic analyses of the Staphylococcaceae family suggest taxonomic reclassification within the genus Staphylococcus. Int J Syst Evol Microbiol 70:5926–5936. https://doi.org/10.1099/ijsem.0.004498
20. Crippa BL, Matos LG, Souza FN, Silva NCC (2024) Non-aureus staphylococci and mammaliicocci (NASM): their role in bovine mastitis and One health. J Dairy Res 91:1–13. https://doi.org/10.1017/S0022029924000165
21. Fatahi-Bafghi M (2021) Characterization of the Rothia spp. and their role in human clinical infections. Infect Genet Evol 88:104686. https://doi.org/10.1016/j.meegid.2020.104686
22. Ruaro A, Andrighetto C, Torriani S, Lombardi A (2013) Biodiversity and characterization of indigenous coagulase-negative staphylococci isolated from raw milk and cheese of North Italy. Food Microbiol 34:106–111. https://doi.org/10.1016/j.fm.2012.11.017
23. Saiki R, Hagi T, Narita T, Kobayashi M, Sasaki K, Asahina Y, Nomura M (2018) Effects of the addition of non-starter lactic acid bacteria on free amino acid production during cheese ripening. Food Sci Technol Res 24:299–309.
24. Kondrotiene K, Zavistanaviciute P, Aksomaitiene J, Novoslavskij A, Malakauskas M (2024) Lactococcus lactis in dairy fermentation—Health-promoting and probiotic properties. Fermentation 10:16. https://doi.org/10.3390/fermentation10010016
25. Perin LM, Belviso S, Dal Bello B, Nero LA, Cocolin L (2017) Technological properties and biogenic amines production by bacteriocinogenic lactococci and enterococci strains isolated from raw goat's milk. J Food Prot 80:151–157. https://doi.org/10.4315/0362-028X.JFP-16-255
Received: March 25, 2026 / Accepted: April 8, 2026 / Published (Online First): April 11, 2026 / Issue Date: June 15, 2026 (Europe/Madrid)
Citation: Ramos GLPA, Silva ECC, Araújo LF, Moraes MS, Nascimento JS. Microbiological characterization of raw goat milk from a small municipality in Brazil: identification of Gram-positive cocci by MALDI-TOF MS. BioNatura Journal: Ibero-American Journal of Biotechnology and Life Sciences. 2026;3(2):1. https://doi.org/10.70099/BJ/2026.03.02.1
Correspondence should be addressed to: gustavoanciens@id.uff.br
Peer Review Information BioNatura Journal thanks the anonymous reviewers for their valuable contribution to the peer-review process. Regional peer-review coordination was conducted under the BioNatura Institutional Publishing Consortium (BIPC). Reviewer selection and assignment were supported via: https://www.reviewercredits.com/
Digital Preservation and Repository. This journal is managed through the Open Journal Systems (OJS) platform. To ensure long-term access, we use the PKP Preservation Network (PKP PN) to digitally preserve all published volumes in a decentralized, secure archive. Furthermore, our repository is integrated with LOCKSS and CLOCKSS, allowing international library networks to create permanent archives for long-term survival.
Publisher Information Published by Clinical Biotec S.L. (Madrid, Spain) as the publisher of record under the BioNatura Institutional Publishing Consortium (BIPC). Places of publication: Madrid (Spain); Tegucigalpa (Honduras); Panama City (Panama). Online ISSN: 3020-7886.
Open Access Statement. All articles published in BioNatura Journal are freely and permanently available online immediately upon publication, without subscription charges or registration barriers.
Copyright and License © 2026 by the authors. This article is published under the terms of the Creative Commons Attribution (CC BY 4.0) license, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. License details: https://creativecommons.org/licenses/by/4.0/
Governance For editorial governance and co-publisher responsibilities, see the BIPC Governance Framework (PDF) at: https://clinicalbiotec.com/bipc
